CDS uses a large variety of techniques situated in the field of proteomics, (epi)genomics and protein biochemistry.
Here are a few examples of equipment in the CDS lab:
Flash photolysis and stopped flow
The reactivity of globins towards gaseous ligands is an important indication for the function these proteins might fulfill in vivo, and depends for a large part on the ligation state of the heme Fe-atom. The ligand binding affinities and kinetics can be determined by flash photolysis and stopped flow.
In flash photolysis experiments, a short laser pulse (5-7 ns) is used to dissociate the bound ligand from the heme iron atom. The use of a laser for sample excitation gives the technique the specificity of single wavelength excitation and nanosecond time resolution. Photo-dissociated ligands first migrate into open spaces within the globin interior and then diffuse back to the distal pocket, where they either rebind to the heme iron atom, a process called geminate recombination, or escape to the solvent. The ligand can then rebind to the heme iron atom from outside the protein matrix, called bimolecular rebinding.
Our lab is equipped with the LP920 laser photolysis system (Edinburg Instruments) coupled to a frequency-doubled Q-switched Nd:YAG laser (Spectra Physics Quanta-ray).
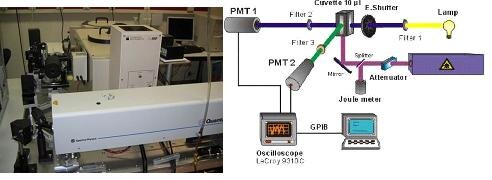
In stopped flow experiments, two reagents are rapidly mixed together and then ‘stopped’ in an observation cell. The sample cell is irradiated with (usually) monochromatic light and as the reaction proceeds the change in the recorded signal, usually a fluorescence signal or the absorbance at a specific wavelength, is recorded as a function of time.
Our lab is equipped with a SX20 Stopped Flow Reaction Analyser from Applied Photophysics.
Microcapillar 2D-LC (Agilent)
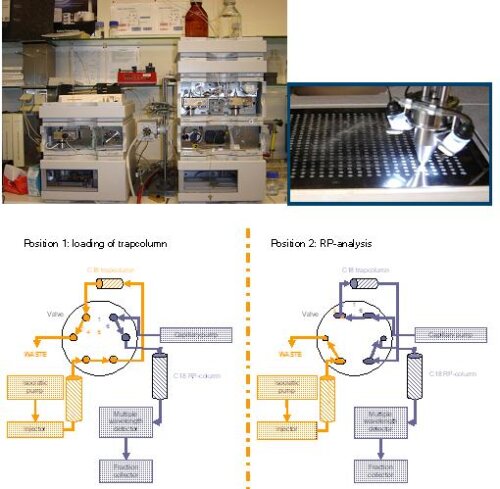
Chromatographic analysis of complex protein samples needs a multidimensial approach as separation of hundreds to thousands of proteins is not possible by one chromatographic step. The lab therefore has setup a configuration whereby the fractions resulting from a Strong Cation Exchange (SCX) microcolumn (800 µm diameter) are each separately run over a Reverse Phase (RP) capillary column (300 µm diameter). Although we usually apply this combination offline, a configuration for online switching from SCX to RP is also available on the lab, allowing for automated and serial 2D-LC chromatography. In this configuration, the SCX column is eluted with stepwise increasing saltplugs so that ‘packets’ of peptides come off from the column, which are subsequently ‘parked’ on a trap column. Valve switching then allows for the peptides to come off from the trap column after which they can be separated on by RP chromatography. Hereafter, a new ‘cycle’ with a saltplug that contains a higher salt concentration can start.
To increase throughput, we installed a configuration with two RP columns so that one column can be used for separation while the other one is washed and equilibrated.The system is also equipped with a spotter device where the eluate from the RP column is automatically spotted on a MALDI target.
QTOF mass spectrometer (Waters)
For the study of protein posttranslational modifications (biotinylation, SUMOylation, methylation, acetylation, coupling to Withaferin), we have available a QTOF mass spectrometer (Waters), preceded by a RP capillary column (500 µm diameter, Agilent). This instrument is most often used for measuring and quantifying modifications on purified proteins.
The instrument is also used for validading proteotypic peptides for their later use in MRM measurements on a triple quadrupole mass spectrometer.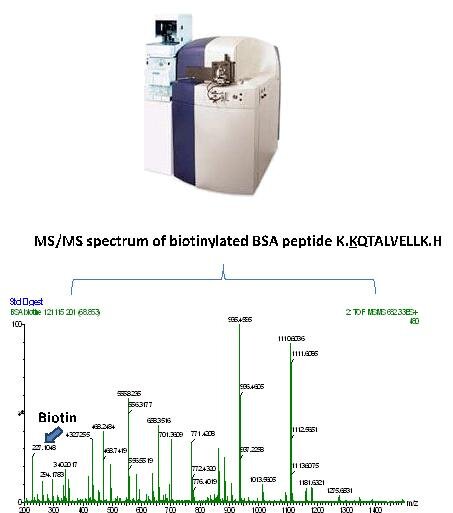
Procise amino acid sequencer
An automated amino acid sequencer uses the chemistry developed by Pehr Edman in the 1950s for the sequential degradation of proteins and peptides to determine the N-terminal amino acid sequence.
The lab is equipped with a Procise protein sequencing system from applied Biosystems.
Pyromark Q24 pyrosequencer
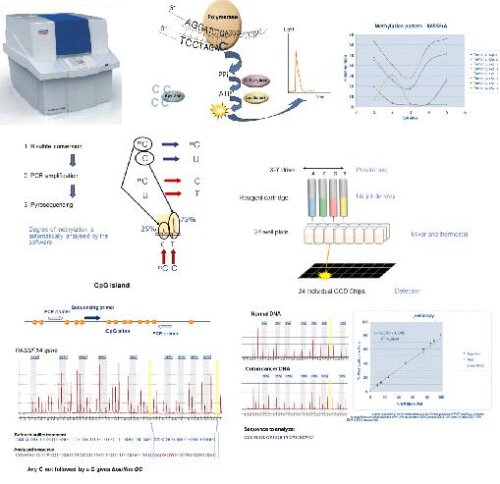
Bisulfite sequencing is the use of bisulfite treatment of DNA to determine its pattern of methylation. DNA methylation was the first discovered epigenetic mark, and remains the most studied. In animals it predominantly involves the addition of a methyl group to the carbon-5 position of cytosine residues of the dinucleotide CpG, and is implicated in repression of transcriptional activity.Treatment of DNA with bisulfite converts cytosine residues to uracil, but leaves 5-methylcytosine residues unaffected. Thus, bisulfite treatment introduces specific changes in the DNA sequence that depend on the methylation status of individual cytosine residues, yielding single- nucleotide resolution information about the methylation status of a segment of DNA.
Following PCR amplification of the region of interest, CpG pyrosequencing is used to determine the bisulfite-converted sequence of specific CpG sites in the region. The ratio of C-to-T at individual sites can be determined quantitatively based on the amount of C and T incorporation during the sequence extension.
PamStation 12
The PamStation 12 is a fully automated instrument designed for processing Pamgene peptide microarrays.